Mutanome.Ai™ Supports Precision Therapy Decisions by Functional Interpretation of Coding and Non-Coding Variants

Mutations, essentially spelling errors in our genetic code occur frequently, but most are promptly corrected by DNA repair mechanisms. A small fraction, however, escapes and becomes permanent changes in the DNA. Fortunately, the vast majority of those mutations are benign (harmless) and do not result in any observable phenotypic changes. However, rare variants/mutations are associated with many human non-heritable and heritable diseases. Genome sequencing has gained momentum as a tool to detect inherited genetic predispositions, and a corpus of accumulated pathogenic variants are driving the development of next generation healthcare, novel drugs and personalized medicines. The mutational profiles of diseases are powerful biomarkers which guide treatment decisions tailored to an individual’s unique genetic makeup.
While mutation detection has become relatively streamlined and cost-effective, reliably identifying disease-relevant pathogenic mutations and predicting their functional impact remains a bottleneck. A large fraction of mutations identified through individual genome sequencing or genome wide association studies (GWAS) are classified as variants of unknown significance (VUS), providing little to no clinically actionable insights. To address this, various computational approaches including machine learning models trained on DNA and protein data has been developed. However, these models still suffer from high false positive/negative rates, limited accuracy, and a lack of contextual biological information. These limitations largely stem from insufficient training data and the inability of single-feature training (focused on either DNA or protein) to capture the full spectrum of pathogenicity. Moreover, accurate, context-specific functional interpretation often requires expert input to identify potential biomarkers or therapeutic targets. This remains a critical capability gap, limiting the speed and precision of therapeutic decision making in clinical settings.
Overview of Mutanome.Ai™
ThinkBio’s Mutanome.Ai™ is an AI powered DNA analysis pipeline to reliably detect and interpret pathogenic mutations at scale for effective clinical decision making. It deploys state of the art DNA and protein foundation models, for variant effect prediction (VEP) and sequence to function evaluation. Although coding mutations have been extensively studied - particularly for identifying therapeutic targets- over 98% of the human genome is non-protein-coding. Mutations in these non-coding regions remain largely unexplored, yet they hold significant potential for uncovering novel biomarkers and treatment targets. Mutanome.Ai™ offer a comprehensive strategy for detecting pathogenic mutations in both the coding and the non-coding sequences.
Mutanome.Ai™ allows users to shortlist pathogenic mutations tailored to their research questions. Once the likely pathogenic coding sequence-alternating mutations are identified, Mutanome.Ai performs a downstream functional analysis assessment to uncover key genes and pathways impacted by the mutations. By integrating a protein-structure model the impact of the alternations on protein-interaction networks are evaluated. This enables the detection of optimal drug targets within the enriched pathways. In addition, mutational profiles could become important indicators of comorbidity conditions in the patient genome.
Furthermore, Mutanome.Ai™ employs a state-of-the art sequence-to-function model, to predict the impact of likely pathogenic mutations on changes in the gene expression level. In our validation study, Mutanome.Ai reliably predicted the gene expression impact of previously reported regulatory mutations in the promoter region of TERT (Telomerase Reverse Transcriptase enzyme), a cancer-driver gene whose expression level and regulatory mutations are used as biomarkers to predict the stability of telomeric regions of chromosomes.
In a second case study we further evaluated a list of single nucleotide variants reported to be associated with breast cancer (GWAS catalog EFO ID: MONDO_0007254) through Mutanome.Ai™ and identified a pathogenic mutation predicted to reduce the expression of a tumour suppressor gene, PHLDA3. Downregulation of PHLDA3 has already been explored as an expression-biomarker for breast cancer and our finding via Mutanome.Ai™ identified a potential cancer-associated mutation as a biomarker. Even though minor allele frequency (MAF) of this mutation is around 0.06 in European population, considerable variation in MAF has been reported in other populations. Therefore, the predictive value of this biomarker should be further assessed in a population-specific context for which Mutanome.Ai offers an ideal evaluation tool. Taken together, these findings strongly demonstrate that Mutanome.Ai is a fast, reliable and a scalable AI-enabled novel biomarker discovery pipeline.
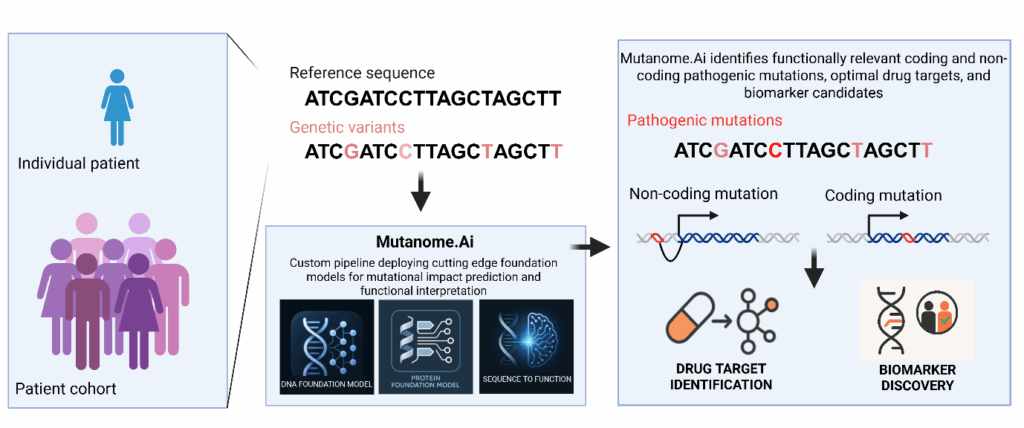
Starting with genetic variants detected from individual patients or cohort data, Mutanome.Ai™ deploys state of the art foundation models and perform prediction quality assessments to identify optimal drug targets and clinically actionable biomarkers.
Mutanome.Ai™ enables faster and effective precision therapy decisions
Mutanome.Ai™ is easily integrated with other omics pipelines such as transcriptome and epigenome to provide additional layers of evidence for making therapy decisions. By integrating Mutanome.Ai™ with ThinkBio’s proprietary knowledge platform, TheraBluePrint™, we provide robust biological support, enriching the short-listed variants with disease-specific insights.
In summary, following are the different use cases of Mutanome.Ai™.
Patient mutation evaluation: Mutanome.Ai™ can perform mutation evaluation for individual patients and detect potential comorbidities.
Mutation annotation and biomarker discovery: Mutanome.Ai™ can annotate mutations detected in large cohort studies including UK Biobank, All of Us, other national genomic initiatives, large consortia, or GWAS studies. It can then perform functional impact prediction to discover novel biomarkers.
Drug target identification: Interaction network of genes with pathogenic mutations in a cancer type can provide actionable insights to clinical labs and pharmacogenomics companies. Also, can identify population specific prioritized treatment targets by evaluating population datasets.
Plug in tool: Mutanome.Ai™ pipeline can be offered as Plug-in tool with large-scale knowledge platforms (TheraBluePrint™ ) or other bioinformatics platforms by Cloud providers (AWS Healthomics, for example).
AI tools are revolutionizing biomedical research and healthcare sector by enabling rapid data processing, integration, and interpretation at an unprecedented scale. Mutanome.Ai™ pipeline represents a fundamental leap forward addressing a critical capability gap in extracting clinically actionable information from genomic datasets.