Cyclin-Dependent Kinases as Key Targets in Cancer Therapy
Cancer arises when cells grow and divide uncontrollably, often due to failures in the normal checkpoints of the cell cycle. The cell cycle is tightly regulated by one of the central regulatory mechanisms involving cyclin-dependent kinases (CDKs), which pair with specific cyclins at defined stages (Figure 1). These CDK–cyclin complexes act like traffic lights, ensuring that cells progress only when conditions are appropriate (1).
Figure 1: The cell cycle: G1 (cell growth and DNA synthesis preparation), S (DNA replication), G2 (preparation for mitosis), and M (mitosis). Cells can also exit the cycle into a resting state called G0.
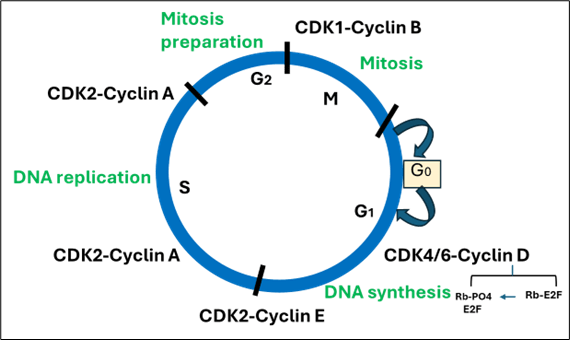
Given their central role in cell-cycle regulation, CDKs have long been considered attractive targets in cancer therapy. Early strategies that broadly inhibited multiple CDKs caused significant toxicity, as these enzymes are highly conserved and essential for normal cell function. This challenge has shifted the field toward a more selective, biology-driven approach (1). One of the most promising points of intervention in the cell cycle is the G1–S transition, where a cell commits to copying its DNA and dividing. This step is tightly controlled by a sequence of CDK–cyclin complexes. CDK4/6, CDK2, and CDK1 are key regulators of cell-cycle progression that act at distinct checkpoints by forming complexes with cyclins and phosphorylating essential cell-cycle proteins. CDK4/6 initiates the phosphorylation process in early G1 by partially inactivating the Rb (Retinoblastoma) protein. CDK2 then completes this phosphorylation, pushing the cell into S phase, where DNA replication occurs. Later, CDK1 controls the critical step of entry into mitosis and is generally considered the most important CDK for canonical cell division across most normal mammalian cell types (Figure 1) (2).
Recent studies indicate that CDK1 and CDK2 exhibit high plasticity and can interact with multiple cyclins (primarily cyclins A, B, D and E), whereas CDK4 and CDK6 have a more limited range, primarily partnering with D-type cyclins. The broader interaction capacity of CDK2, and its frequent overactivation in various cancers, underscores its appeal as a therapeutic target. Moreover, CDK2 can form non-canonical complexes with cyclin D to bypass the G1 arrest caused by CDK4/6 inhibitors, emphasizing its central role in sustaining cancer cell proliferation. This suggests that CDK4/6 inhibition alone may not be insufficient due to this bypass mechanism, and that combinatorial strategies targeting both CDK2 and CDK4/6 could be more effective (3,4,5).
Since CDK1 plays a vital role in all dividing cells, including healthy tissues, even partial inhibition causes significant toxicity. In contrast, CDK2 plays a more supportive role at the G1–S boundary. Normal cells can often compensate for the loss of CDK2 by using CDK1, but many cancer cells cannot, particularly those with cyclin E amplification prefers to bind with CDK2. In tumours, CDK2 is frequently overactive due to elevated cyclin E levels, oncogenic signalling, or loss of natural CDK inhibitors. As a result, cancer cells become more dependent on CDK2 than normal cells (2,3).
Although CDKs share a conserved ATP-binding site, small structural differences between CDKs particularly in the hinge region and nearby pockets allow the development of inhibitors that selectively target CDK2. This has enabled new therapeutic approaches, including selective ATP-competitive, covalent, and emerging allosteric inhibitors. CDK2 has become an important cancer drug target because it plays a central role in controlling the G1–S transition and S-phase entry. It is frequently dysregulated in cancer, often through Cyclin E1 amplification, and can form non-canonical complexes that bypass the cell-cycle arrest caused by CDK4/6 inhibitors. Selective inhibition of CDK2 will effectively block Rb phosphorylation suppressing E2F-driven transcription, leading to reduced cancer cell proliferation. Targeting CDK2 is especially powerful in combination with CDK4/6 inhibition offers a more effective strategy for modern anticancer therapy. Establishing the non-overlapping role of CDK2 and CDK1 in regulation of cell cycle took the scientific community decades of experimentation and accumulation of data. Recently, AI-based methods have begun to integrate multi-omics and pathway data to refine our understanding of CDK1, CDK2, and CDK4/6, specifically defining their roles in cell-cycle control. These approaches streamline target prioritization by predicting pathway dependencies, druggability, and potential resistance mechanisms. The use of AI tools can accelerate this process significantly, revealing novel biology and bringing unappreciated drug targets into the druggable arena.
At ThinkBio.Ai®, we employ an integrative systems biology approach combined with AI-integrated protein networks to discover druggable gene targets and their associated protein networks. By analysing signalling pathways, protein–protein interactions, and structural features across multiple biological scales, this strategy enables the rational prioritization of targets that play central roles in disease-relevant processes. This integrated framework supports the discovery and development of selective, mechanism-driven therapeutics with improved efficacy and safety profiles.
References:
1. Malumbres M, Barbacid M. Cell cycle, CDKs and cancer: a changing paradigm. Nat Rev Cancer. 2009;9(3):153–166. doi:10.1038/nrc2602.
2. Rubin SM, Sage J, Skotheim JM. Emerging strategies to inhibit the G1/S transition for cancer therapy. Cancer Res. 2026; Epub ahead of print. doi:10.1158/0008-5472.CAN-25-0916.
3. Kumarasamy V, Nambiar R, et al. CDK/cyclin dependencies define extreme cancer cell-cycle heterogeneity and collateral vulnerabilities. Cell Rep. 2022;38(10):110448. doi:10.1016/j.celrep.2022.110448.
4. Putta S, Villegas CA, Rubin SM. Differences in binding affinity among cell-cycle CDK and cyclin pairs. J Mol Biol. 2025;437(5):168952. doi:10.1016/j.jmb.2025.168952.
5. Knudsen ES, Witkiewicz AK, Sanidas I, Rubin SM. Targeting CDK2 for cancer therapy. Cell Rep. 2025;44(8):116140. doi:10.1016/j.celrep.2025.116140.